| Kidney Res Clin Pract > Epub ahead of print |
Abstract
Background
Methods
Results
Supplementary Materials
Notes
Funding
This work was supported by the Basic Science Research Program (NRF-2022R1F1A1071128 to Samel Park and 2021R1C1C1007810 to Jong-Seok Moon) from the National Research Foundation of Korea.
Data sharing statement
The data presented in this study was obtained from the Korean Genome-wide Epidemiology Study (KoGES). It could be requested on the website (https://biobank.nih.go.kr/).
Authors’ contributions
Conceptualization, Formal analysis, Visualization: SP
Data curation: DSK, NJC
Funding acquisition, Project administration: JSM, SP
Investigation: DJL, SP
Methodology: DKS, YSL
Resources: DJL, NJC
Software: DJL, DSK
Supervision: JSM, YSL, HWG, EYL
Writing–original draft: DJL, SP
Writing–review & editing: DJL, SP
All authors read and approved the final manuscript.
Acknowledgments
Figure 1.
Circos plot for GWAS meta-analysis.
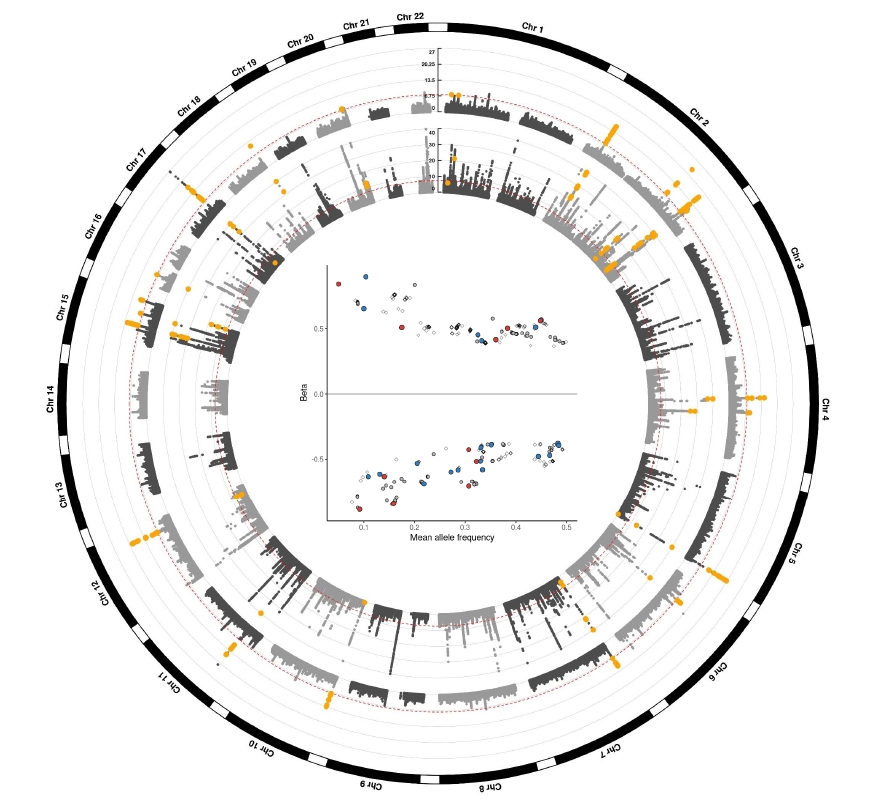
Figure 2.
Independent significant SNPs validated by at least one biomarker, log-BUN or eGFRcysC.
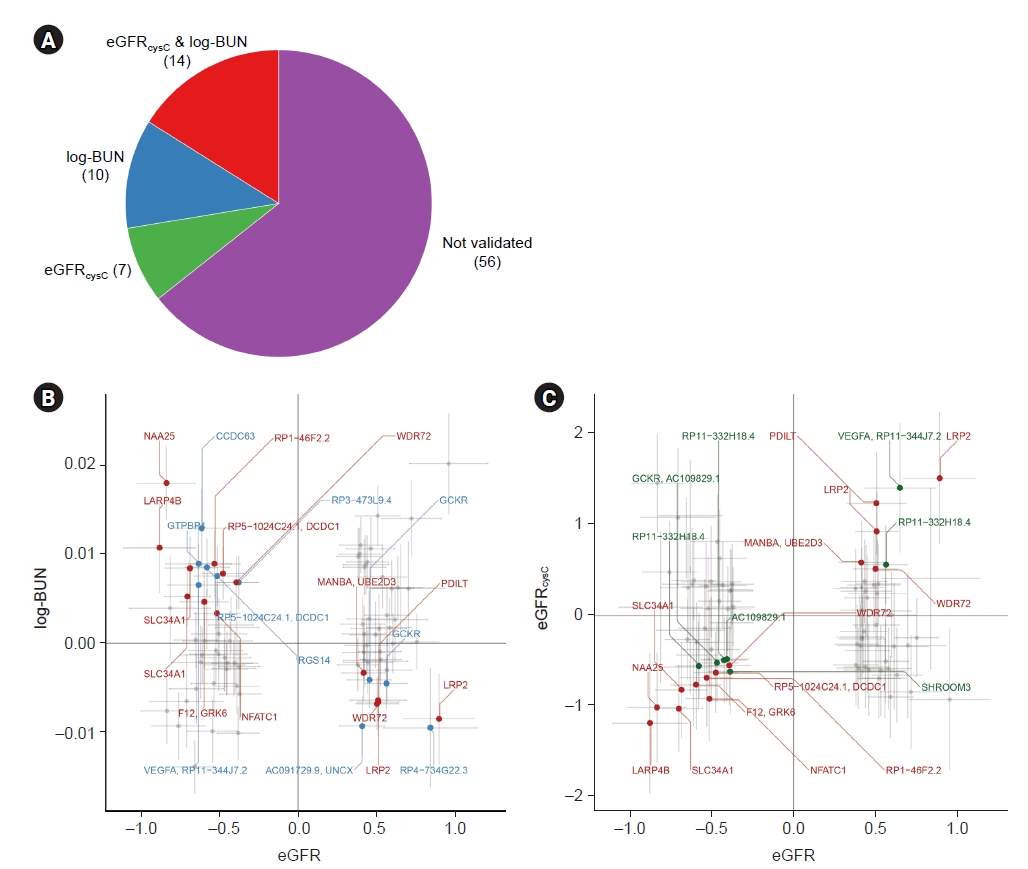
Figure 3.
Scatter plot for concordance between studies.
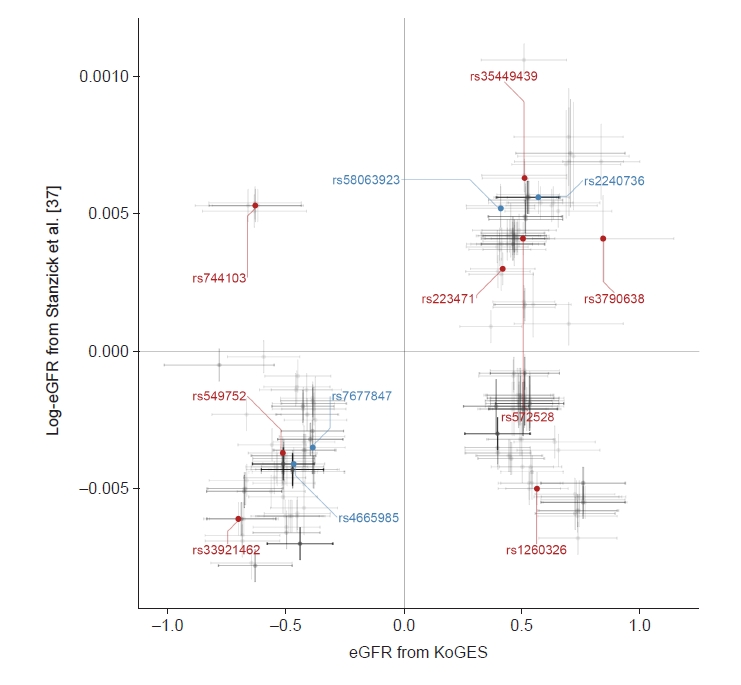
Figure 4.
MAGMA gene analysis and pathway analysis based on significant genes.
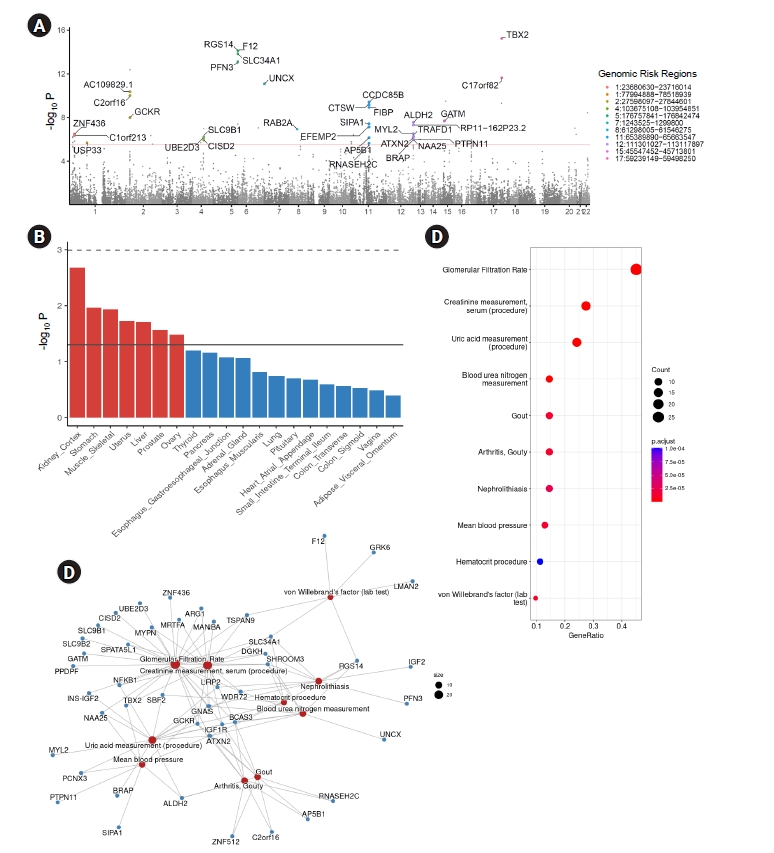
Figure 5.
Heritability enrichment in tissues from GTEx and Franke using LDSC-SEG.
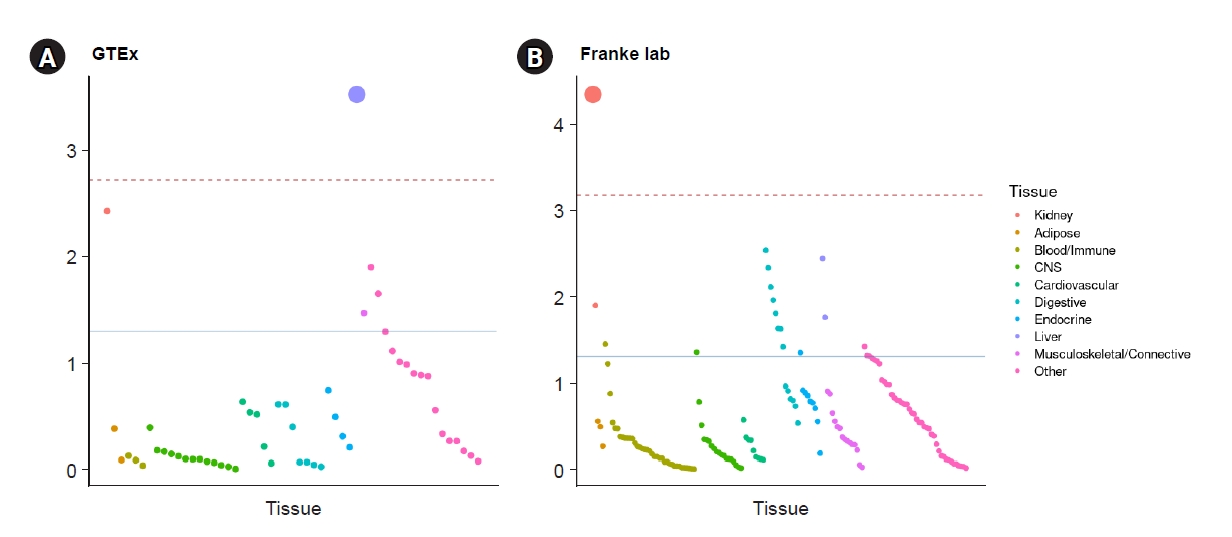
Table 1.
Data are expressed as number only, median (interquartile range ), or number (%).
There were some missing values: 1) 6 for waist circumference, 24 for glucose level, 1 for HbA1c, and 3 for ALT in KARE cohort; 2) 2 for DBP, 7 for waist circumference, 5 for hemoglobin, 5 for HbA1c, 1 for AST, 29 for ALT in HEXA cohort; 3) 14 for waist circumference, 475 for HbA1c, 5 for ALT, and 80 for cystatin C in CAVAS cohort.
ALT, alanine transferase; AST, aspartate transferase; BUN, blood urine nitrogen; BMI, body mass index; CAVAS, Cardiovascular Disease Association Study; DBP, diastolic blood pressure; eGFR, estimated glomerular filtration rate calculated using serum creatinine by CKD-EPI equation; eGFRcysC, estimated glomerular filtration rate calculated using serum cystatin C by CKD-EPI equation; HbA1c, glycated hemoglobin; HEXA, Health Examinee; KARE, Korean Association Resource; SBP, systolic blood pressure.
Table 2.
| CHR | Genomic risk region | rsID | Position (b37) | EA/NEA (EAF) | BETA | p-value | Cond P | Mapped gene lista |
|---|---|---|---|---|---|---|---|---|
| 1 | 1:10,702,266–10,718,377 | rs3790638 | 10,707,812 | A/G (0.05) | 0.84 | 3.6 × 10–8 | RP4-734G22.3 | |
| 2 | 2:27,598,097–27,844,601 | rs1260326b | 27,730,940 | C/T (0.45) | 0.56 | 9.6 × 10–17 | – | GCKR |
| rs4665985c | 27,753,878 | A/C (0.47) | –0.47 | 2.2 × 10–12 | 2.4 × 10–37 | GCKR, AC109829.1 | ||
| rs62141288 | 27,783,198 | A/G (0.33) | –0.41 | 7.6 × 10–9 | 7.6 × 10–25 | AC109829.1 | ||
| 2 | 2:170,165,283–170,206,062 | rs77366165 | 170,170,804 | A/G (0.10) | 0.90 | 3.9 × 10–15 | LRP2 | |
| 4 | 4:77,363,639–77,414,988 | rs7677847 | 77,364,126 | A/G (0.35) | –0.39 | 2.6 × 10–8 | SHROOM3 | |
| 4 | 4:103,675,108–103,954,851 | rs223471b | 103,698,786 | C/G (0.36) | 0.42 | 2.9 × 10–9 | MANBA, UBE2D3 | |
| 5 | 5:176,757,841–176,842,474 | rs33921462b | 176,814,656 | A/G (0.31) | –0.70 | 8.0 × 10–21 | SLC34A1 | |
| 6 | 6:43,804,571–43,829,941 | rs744103b | 43,805,362 | T/A (0.14) | –0.63 | 2.3 × 10–10 | – | VEGFA, RP11-344J7.2 |
| rs35578578 | 43,810,526 | G/GC (0.10) | 0.65 | 4.5 × 10–9 | 6.6 × 10–25 | VEGFA, RP11-344J7.2 | ||
| 7 | 7:1,243,525–1,299,800 | rs58063923c | 1,270,738 | T/C (0.33) | 0.41 | 3.7 × 10–8 | AC091729.9, UNCX | |
| 10 | 10:847,688–1,081,293 | rs17159964 | 913,064 | T/G (0.09) | –0.88 | 3.2 × 10–14 | LARP4B | |
| 11 | 11:30,749,090–30,777,790 | rs56870952 | 30,750,092 | T/TACAAAACAAA (0.33) | –0.58 | 2.9 × 10–16 | RP5-1024C24.1, DCDC1 | |
| 12 | 12:111,301,027–113,117,897 | rs11066132b | 112,468,206 | T/C (0.16) | –0.84 | 9.0 × 10–20 | NAA25 | |
| 15 | 15:53,882,419–54,006,275 | rs572528 | 53,972,484 | A/G (0.38) | 0.50 | 2.9 × 10–13 | WDR72 | |
| 16 | 16:20,383,049–20,407,196 | rs35449439 | 20,385,182 | C/G (0.18) | 0.51 | 3.0 × 10–8 | PDILT | |
| 17 | 17:59,239,149–59,498,250 | rs2240736b | 59,485,393 | T/C (0.45) | 0.57 | 2.8 × 10–17 | RP11-332H18.4 | |
| 18 | 18:77,156,103–77,160,235 | rs549752b | 77,158,225 | A/G (0.32) | –0.52 | 3.8 × 10–12 | NFATC1 |
CHR, chromosome; EA, effect allele; EAF, effect allele frequency; NEA, non-effect allele; rsID, reference SNP cluster ID; SNP, single nucleotide polymorphism.
References
- TOOLS
-
METRICS

-
- 0 Crossref
- 0 Scopus
- 1,046 View
- 40 Download
- ORCID iDs
-
Dong-Jin Lee

https://orcid.org/0009-0000-7265-8299Jong-Seok Moon

https://orcid.org/0000-0002-2537-7854Dae Kwon Song

https://orcid.org/0000-0002-5086-2093Yong Seok Lee

https://orcid.org/0000-0002-8687-589XDong-Sub Kim

https://orcid.org/0009-0007-0854-9750Nam-Jun Cho

https://orcid.org/0000-0002-9053-0499Hyo-Wook Gil

https://orcid.org/0000-0003-2550-2739Eun Young Lee

https://orcid.org/0000-0002-4513-9888Samel Park

https://orcid.org/0000-0002-5717-0743 - Related articles



 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Supplement table 1
Supplement table 1 Print
Print