Manifestation of rs1888747 polymorphisms in the FRMD3 gene in diabetic kidney disease and diabetic retinopathy in type 2 diabetes patients
Article information
Abstract
Background
FRMD3 polymorphisms has suggested that they could be an alternative test to differentiate diabetic nephropathy (DN) from nondiabetic renal disease (NDRD) in type 2 diabetes mellitus (DM) patients. This study was performed to investigate the relationship between the FRMD3 gene and clinical characteristics of DN.
Methods
Patients who already had renal pathologic results were tested for FRMD3 polymorphisms. The subjects were classified into three groups; DN with diabetic retinopathy (DR), DN without DR, and DM with NDRD. FRMD3 polymorphisms were analyzed in each group.
Results
The prevalence of GG, CG, and CC was 44.4%, 42.2%, and 13.3% respectively. There was no significant difference in clinical parameters, which consisted of disease duration, proteinuria, and complications in DN with or without DR and DM with NDRD. The G allele was mainly found in DN with DR patients (50.8%) whereas the C allele was found in DM with NDRD patients (43.5%) (p = 0.02). There was a significant association between the CC genotype in NDRD when compared to GG (p = 0.001). In addition, the C allele was 2.10-fold more often associated with NDRD than the G allele (p = 0.03). The CC genotype was correlated with risk for NDRD than the GG and GC genotypes, with odds ratios of 6.89 and 4.91, respectively (p = 0.02).
Conclusion
C allele presentation, especially homozygous CC, was associated with NDRD pathology in patients with overt proteinuria. Hence, kidney biopsy is suggested in those with the C allele or homozygous CC genotype, regardless of retinopathy manifestations.
Introduction
Diabetic kidney disease or diabetic nephropathy (DN) are two of the most common complications in diabetic patients. Approximately 20% to 40% of diabetic patients develop diabetic kidney disease, which can progress to chronic kidney disease [1–3]. Its pathophysiology is similar to the progression of diabetic retinopathy (DR). Recent studies showed that both metabolic and hemodynamic stimuli could activate key intracellular signaling pathways along with transcription factors that contribute to microvascular damage in both glomeruli and the retina [1,4,5].
The diagnosis of DN can be made when patients manifest persistent proteinuria (exceed 500 mg within 24 hours) and hypertension along with a progressive decline in renal function [3,6]. This condition is usually preceded by the stage of microalbuminuria [7]. Without intervention, diabetic patients with microalbuminuria progress to proteinuria and then overt DN [4,8]. However, there are reports of nondiabetic renal disease (NDRD) that are isolated or superimposed on DN, such as immunoglobulin (Ig) A nephropathy, minimal change disease, and idiopathic focal segmental glomerulosclerosis.
The prevalence of NDRD in diabetic patients confirmed by tissue pathology reports ranges from 10% to 85% [9–12]. This wide prevalence range may result from selection criteria, biopsy threshold, or sample size [13]. Due to its indolent nature, chronic kidney disease diagnosed after DR is typically presumed to be DN [5]. However, diabetic patients with high amounts of proteinuria without DR are recommended for a kidney biopsy to rule out other glomerular diseases [13–16]. Because frequent complications from kidney biopsy often occur, many patients may refuse to receive one. Many nephrologists are also reluctant to perform renal biopsy on diabetic patients due to its potential complications, such as hematoma, arterial embolization, and even necessity of nephrectomy. Additionally, many primary hospitals lack the facilities necessary for kidney biopsy. Therefore, nephrologists must provide a diagnosis based on the patients’ clinical manifestations and laboratory results [3,8,10].
Novel studies have observed that the FRMD3 genotype in the rs1888747 position most likely correlates with diabetic kidney disease. The prominent gene expression can be narrowed down to three alleles. First, the CC allele was observed to be a protective gene for diabetic kidney disease, suggesting other glomerular pathology. Second, GC and GG were both progressive alleles which were predisposing factors favoring diabetic kidney disease. Hence, identifying single nucleotide polymorphisms (SNPs) of rs1888747 in the FRMD3 gene from blood cells could potentially serve as a less invasive procedure compared to kidney biopsy [17–23].
The objective of this research was to study the association between SNP expression in the FRMD3 gene and diabetic kidney disease and DR. Furthermore, we attempted to create a diagnostic guideline for the necessity of kidney biopsy to confirm the presence of diabetic kidney disease.
Methods
Clinical data collection
This was a correlation study between gene polymorphisms and renal pathology in patients with type 2 diabetes mellitus (DM) who had greater than 500 mg/day of urine protein at Her Royal Highness Princess Maha Chakri Sirindhorn Medical Center (MSMC), Thailand between March 13, 2019 and March 12, 2020. All patients voluntarily signed informed consent before participating in the study. The protocol was approved by the Human Research Ethics Board of Srinakharinwirot University (No. SWUEC/E-406/2561) in compliance with Declaration of Helsinki in 1964 and its amendments. Data was collected from type 2 DM patients who were older than 18 years of age. We collected details about their history and diagnosis of diabetes as well as its duration, DR and other complications such as cardio/cerebrovascular disease, peripheral vascular disease, DN, and diabetic neuropathy from medical records. Blood pressure, serum creatinine, serum glucose, hemoglobin A1c (HbA1C), and proteinuria were examined on the day of the hospital visit. The FRMD3 genotype was also tested at the same visit. Four weeks later, a renal biopsy was performed to determine the cause of proteinuria. We excluded patients with uncertain biopsy results, those with mixed pathological results, and inadequate biopsy tissue.
Patients who were diagnosed with DN had to have class II pathological results according to the Pathologic Classification of Diabetic Nephropathy 2010 [24]. Class II in this classification was defined as diffuse diabetic glomerulosclerosis mixed with an increase in extracellular material in the mesangium such that the width of the interspace exceeded two mesangial cell nuclei in at least two glomerular lobules [24]. Meanwhile, DR was evaluated and diagnosed according to the Early Treatment Diabetic Retinopathy Study (ETDRS) report number 7, by showing at least one microaneurysm with soft exudates, cotton-wool spots with venous beading, or definitely-present intraretinal microvascular abnormalities [25].
Clinical and laboratory evaluation
Serum glucose was analyzed by hexokinase/glucose-6-phosphate dehydrogenase. Serum creatinine and HbA1C levels were measured by the enzymatic method. Proteinuria levels were analyzed with the urine protein-creatinine ratio, which was analyzed by the benzethonium chloride and enzymatic method with the ARCHITERT C8000 (Abbott, Abbott Park, IL, USA).
DNA isolation and genotyping
Blood samples were tested for the FRMD3 genotype during routine diabetic follow-up. DNA was extracted from patients’ blood leukocytes and was analyzed for genotyping by real-time polymerase chain reaction (RT-PCR). We studied the rs1888747 SNPs which were associated with DN [18]. DNA was extracted from peripheral blood leukocytes with a QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommendations. SNPs (rs1888747) were genotyped using primer probes contained in the human TaqMan SNP Genotyping Assays (Applied Biosystems, Waltham, MA, USA). RT-PCR reactions were conducted in 96-well plates, in 20-µL total reaction volume using 20 ng of genomic DNA, TaqMan GTXpress Master Mix (2×; Applied Biosystems), and TaqMan SNP Genotyping Assays for rs1888747 (20×). Plates were positioned in an RT-PCR thermal cycler (QuantStudio 5 RT-PCR System; Applied Biosystems) and enzyme activation was performed for 20 seconds at 95°C, followed by 45 cycles at 95°C for 3 seconds and 60°C for 30 seconds.
Patient allocation
FRMD3 gene polymorphisms were analyzed as homozygous GG, heterozygous GC, and homozygous CC in accordance to DN with DR, DN without DR, and DM with NDRD groups, respectively.
Statistical analysis
The researchers employed the IBM SPSS version 23.0 (IBM Corp., Armonk, NY, USA). Through descriptive statistics, data collection was arranged into groups in terms of both frequency and percentage. The p-value and standard deviation were compared for continuous data by using the Student t-test. The chi-square test was used to reveal the relationship between continuous and categorical variables. The Spearman correlation coefficient was used to assess the correlation of selection algorithms with continuous and ordinal variables. The p-values of ≤0.05 were considered statistically significant.
Sample size calculation was based on the hypothesis that the ratio of DN to NDRD was 1:1. Appropriate sample size was calculated by setting the statistical power (1-β) level at 0.80 with α error probability at 0.05 [26]. Including the consideration of DN and NDRD incidence in type 2 DM patients, the calculated applicable sample size was 82 (in this study, n = 90).
Results
Data were gathered from 90 type 2 DM patients, and 37.8% of these had end-stage renal disease, requiring 2 to 3 hemodialysis sessions per week. Each patient also underwent DR examination with an ophthalmologist and diabetic neuropathy examination with monofilament testing within 4 weeks after the genotype was confirmed. This allowed accuracy when correlating genotype detection and microvascular complications of type 2 DM.
Demographic data
Of the 90 subjects, 40 (44.4%) were female. The mean age was 61.7 ± 13.1 years old, and the mean duration of DM was 13.9 ± 9.4 years. As for type 2 DM complications, 45 patients (50.0%) had diabetes retinopathy, 52 patients (57.8%) had diabetic neuropathy, and 29 patients (32.2%) had experienced macrovascular complications (cardio/cerebrovascular accident or peripheral vascular disease). As for renal complications, the mean urine protein was 3.43 g/day (range, 0.53–23.80 g/day), and the results from renal biopsies confirmed that 64 cases (71.1%) were DN and the remaining 26 (28.9%) were NDRD, including minimal change disease, focal segmental glomerulosclerosis, and IgA nephropathy. In addition, we also found that patients with retinopathy (n = 45) manifested renal pathology consistent with DN in 40 of 45 patients (88.9%, p < 0.001). However, 22 of 45 patients (48.9%) without retinopathy also showed renal pathology consistent with DN.
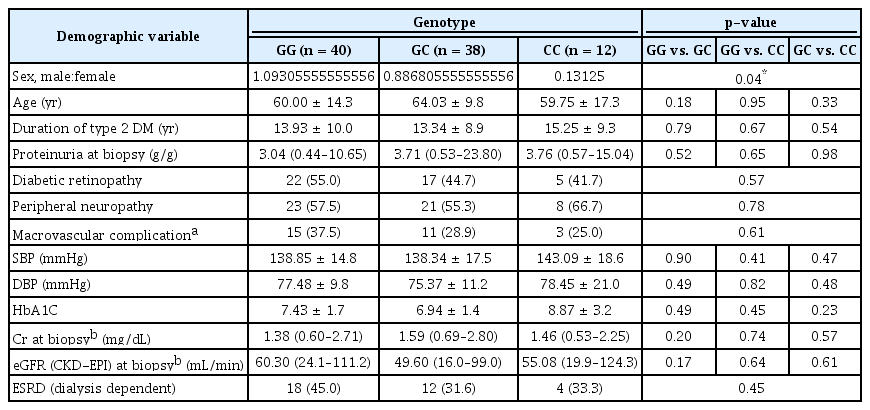
Patients were then categorized into GG, GC, and CC groups according to the rs1888747 genotypic characteristic, and the number of patients in each category was 40 (44.4%), 38 (42.2%), and 12 (13.3%), respectively. With respect to clinical manifestations, as shown in Table 1, there were fewer male subjects in the homozygous CC group (p = 0.04). However, no other significant clinical parameters differed significantly, including years of disease, amount of proteinuria, blood pressure, or rate and severity of microvascular and macrovascular complications. In addition, there was no difference in the amount of proteinuria in patients with DN with or without DR and DM with NDRD (Fig. 1). Therefore, only polymorphism variables were analyzed further in multivariable regression models (Table 1, 2).
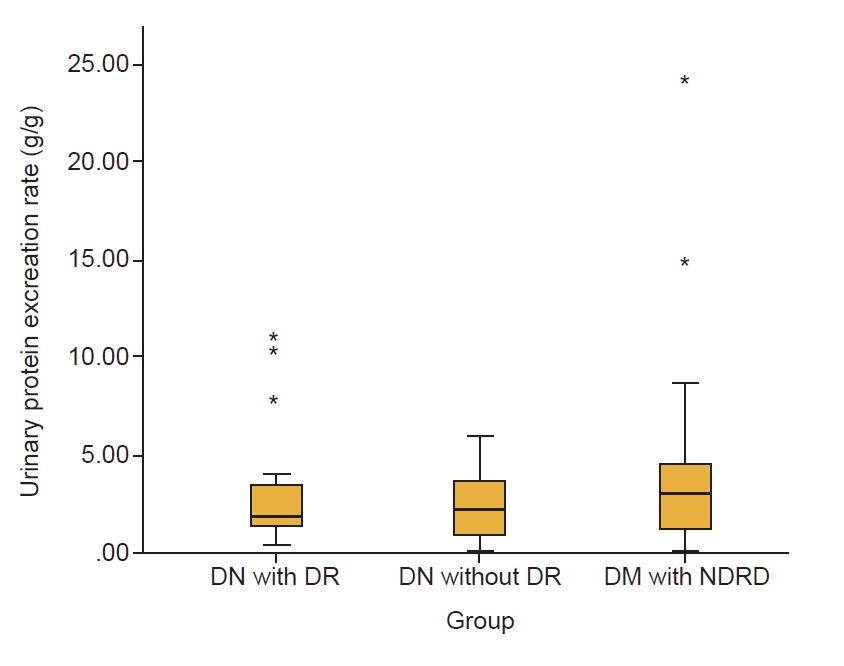
Comparison of proteinuria levels between clinical groups (excluding end-stage renal disease).
Diabetic nephropathy (DN) with diabetic retinopathy (DR) (n = 19), 3.18 g/day (range, 0.44–11.36 g/day); DN without DR (n = 16), 2.46 g/day (range, 0.13–5.96); and type 2 diabetes mellitus (DM) with nondiabetic renal disease (NDRD) (n = 19), 4.75 g/day (range, 0.23–23.80 g/day). DN with DR vs. DN without DR, p = 0.60; DN with DR vs. DM with NDRD, p = 0.24; DN without DR vs. DM with NDRD, p = 0.1, respectively. All outliers were patients with the GG polymorphism (asterisks).
The correlation between the rs1888747 single nucleotide polymorphisms polymorphism of the FRMD3 gene with overt proteinuria and diabetic retinopathy
The distribution pattern of alleles and genotypes are reported in Fig. 2 and 3. The G allele was found in DN with DR, DN without DR, and DM with NDRD 50.8%, 24.6%, and 24.6% of the time, respectively. Meanwhile, the C allele was found in 32.3%, 24.2%, and 43.5% of cases, respectively. The results indicated that the G allele was found mainly in patients with DN whereas the C allele was found in patients with NDRD (p = 0.02) (Fig. 2). Moreover, the CC polymorphism was found significantly more often in patients with NDRD when compared to the GG polymorphism (p = 0.04) (Fig. 3).
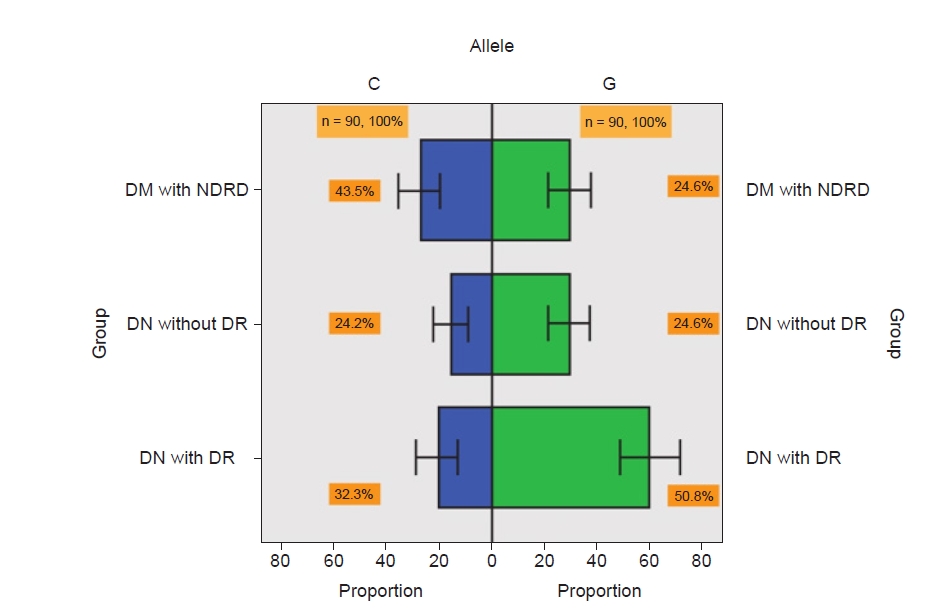
Distribution of G and C alleles in diabetic nephropathy (DN) with diabetic retinopathy (DR), DN without DR, and type 2 diabetes mellitus (DM) with nondiabetic renal disease (NDRD).
The proportion of the G allele was 50.8%, 24.6%, and 24.6%, respectively, in each group. Meanwhile, the proportion of the C allele was 32.3%, 24.2%, and 43.5%, respectively (p = 0.02). The error bars mean 95% confidence intervals.
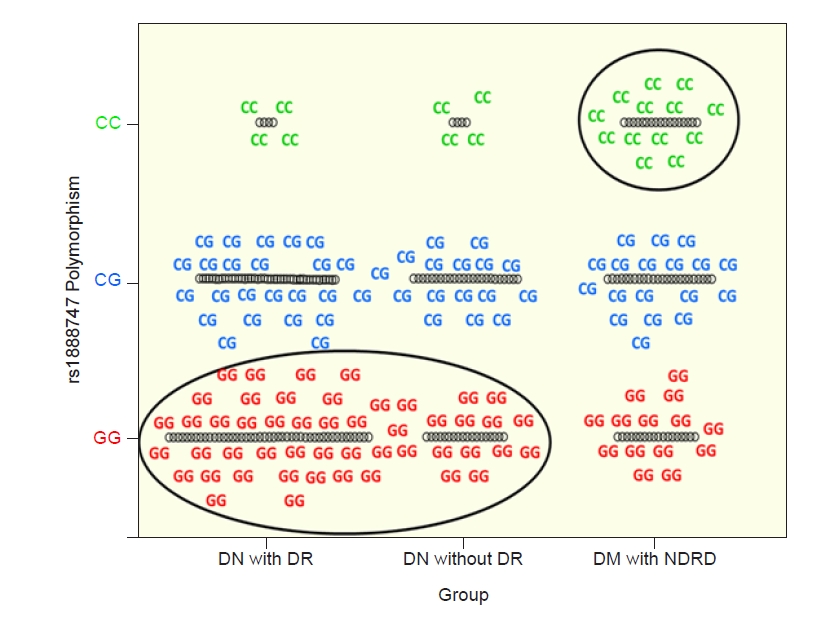
Distribution of genotypes in diabetic nephropathy (DN) with diabetic retinopathy (DR), DN without DR, and type 2 diabetes mellitus (DM) with nondiabetic renal disease (NDRD).
The distribution of the GG polymorphism (red) in each clinical group was 55.0%, 22.5%, and 22.5%, and the distribution of the CG polymorphism (blue) was 42.1%, 28.9%, and 28.9%, respectively. In addition, the distribution of the CC polymorphism (green) in each clinical group was 16.7%, 16.7%, and 66.7%, respectively. The circle indicates that the GG polymorphism manifested in DN with or without DR. Meanwhile, the CC polymorphism was mainly found in the NDRD group (p = 0.04).
The risk of nondiabetic renal disease in diabetic patients based on the rs1888747 single nucleotide polymorphisms polymorphism in the FRMD3 gene
The association between genotype and renal pathology in diabetic patients with overt proteinuria and DR is mentioned in Fig. 2 and 3. These results indicate that patients with the C allele and genotype CC were presumed to have NDRD. Hence, subgroup analysis and logistic regression of patients with and without DR were performed to identify the predicted risk of developing NDRD in diabetic patients, as shown in Table 2. The C allele was 2.10 times more likely to be associated with NDRD, compared to the G allele (p = 0.03). Furthermore, the CC genotype increased the risk of NDRD compared to other genotypes (omnibus test of model coefficients p = 0.02); this genotype carried a 6.89-fold risk compared to the genotype GG by multivariate analysis (p = 0.02). However, patients with the GC genotype were 4.91-fold more likely to have NDRD when compared to those with the GG genotype, although without statistical significance by multivariate analysis.
In the subgroup analysis of patients with and without DR, each group contained 45 participants. Using the power of the test (Zβ) to find the reliability of the subgroup analysis (effect size), it was found that the power was more than 80% in both groups (94% in both groups), which indicated high reliability of results from subgroup analysis. We found that patients with DR who had the C allele had more than a 7-fold risk of NDRD in addition to DN when compared to those carrying the G allele (p = 0.01). This could be interpreted as a 33-fold risk when comparing CC to GG (p = 0.01). Using multivariate analysis, we found that once patients with DR as their only diabetic complication developed proteinuria, it could not promptly be concluded that DN was the pathology responsible for the protein leakage, especially in patients with the CC polymorphism. In contrast, in type 2 DM patients who had proteinuria but did not have DR, there was no statistical difference among polymorphisms that increased the risk of NDRD.
Discussion
The diagnosis of DN predominantly relies on clinical judgment. Important findings contributing to the diagnosis of DN in type 2 DM patients with abnormal proteinuria usually include 1) a diagnosis of type 2 DM for longer than 10 years; 2) the presence of DR; 3) the absence of urine sediment; and lastly, 4) negative viral serologies and immunologic tests for systemic lupus erythematosus [15]. If clinical clues suggesting a different diagnosis from DN are present, renal biopsy should be considered to avoid missing a diagnosis of NDRD. Literature indicates that diabetic patients with NDRD showed significant improvements in proteinuria and renal function following treatment with immunosuppressive agents [15,27]. Patients with NDRD that has similar clinical features to DN, such as primary focal segmental glomerulonephritis, membranous nephropathy, or IgM nephropathy, may not receive a renal biopsy. This may result in a misdiagnosis, leading to a loss of opportunity for treatment. Thus, the development of a tool to distinguish between NDRD and DN in type 2 DM patients with proteinuria is of vital clinical importance, especially in regards to disease management and prognosis.
This study demonstrated that clinical parameters such as glucose level, urine protein level, and the presence of diabetic complications such as retinopathy in diabetic patients cannot distinguish DN from other glomerular diseases. Furthermore, nearly half of diabetic patients without retinopathy also manifested renal pathology consistent with DN.
In previous studies, diabetic patients who had over 10 years of diabetes with overt proteinuria and no DR were often perceived as having other glomerular pathology other than DN. Data showed that in type 2 DM patients, DN tended to develop with DR [5,13,14,16,28,29], suggesting that patients diagnosed with overt proteinuria alone likely had other glomerular diseases rather than DN since there was no evidence of DR. This subset of patients was recommended to undergo kidney biopsy to further evaluate the correct clinicopathologic cause [3,4,7,8,10,15,16].
We have shown that the most frequent SNP polymorphism of the FRMD3 gene at rs1888747 location was GG, then CG, and then CC. Moreover, the G allele and homozygous GG specifically appeared to be more dominant in DN, while C allele and homozygous CC were less common. This observation was similar to previous studies [17–23] which found that expression of the C allele and CC genotype resulted in the prevention of DN. In our report, however, expression of the C allele and CC genotype in patients without retinopathy did not lead to a significant increase in DN when compared to the GG and CG genotypes. On the other hand, patients with DR who expressed the CC polymorphism might have a 33-fold increased risk for NDRD compared to those with the GG polymorphism. These results demonstrated that the occurrence of the C allele and homozygous CC genotype increases the risk of diabetic patients acquiring NDRD as compared to DN.
Based on the results mentioned above, the clinical management of diabetic patients should correspond with the given evidence. A close observation approach is reasonable in patients diagnosed with overt proteinuria without retinopathy (assuming that the renal pathology is DN). However, this approach requires acknowledgment that this group of patients exhibits SNP polymorphisms of the FRMD3 gene with a homozygous GG phenotype to further support the diagnosis. However, if the homozygous CC genotype is present despite having DR, a kidney biopsy is advised in order to identify the differential diagnosis of other glomerular diseases.
Although the CC genotype was associated with NDRD, there were no statistically significant differences between the GG, GC, and CC genotypes in other diabetic complications such as retinopathy, neuropathy, and macrovascular complications. We hypothesized that this was because of bio-differences. The bio-differences are the differences between genotype and phenotype; even though each target tissue expressed the same gene (rs1888747), they create diabetic complications differently due to differences in the hemodynamic system and the exposure to metabolic triggers. For example, the kidney receives an overwhelming portion of the total cardiac output compared to other organs and also has the ability to concentrate solutes. Therefore, the kidneys accumulate metabolites, increasing their effect on the kidney. Therefore, there can be varied outcomes even though different tissue may express the same genes [30–32]. This is a compelling field that has not been thoroughly studied and is an excellent objective for future research. Another limitation of this study is its sample size. Hence, we cannot convincingly show that NDRD can be diagnosed without a renal biopsy based on the predominant pattern of the C allele, as the reason C allele is predominantly found in NDRD may be due to the small proportion of the G allele as a risk factor for DN.
Nonetheless, there are a handful of patients who refuse to undergo kidney biopsy since it is an invasive procedure that may lead to life-threatening complications including intraabdominal hemorrhage, kidney failure, and death. This study supports close monitoring under the assumption that DN develops in patients with the homozygous GG polymorphism in the FRMD3 gene. However, a kidney biopsy is suggested in patients who test positive for the CC or CG phenotypes in order to diagnose other glomerular diseases. The limitation of this study was the absence of qualitative measurements of gene expression levels from each phenotype. Further research should identify the predictive power of these genes and the occurrence of the disease via a receiver operating characteristic curve analysis.
In conclusion, this study demonstrated that in patients diagnosed with type 2 DM, parameters such as glomerular filtration rate, level of proteinuria, and presence of DR were unable to distinguish DN from other glomerular diseases. Meanwhile, the C allele may manifest a certain phenotype that has a renoprotective effect against DN. Therefore, we can imply that there may be underlying glomerular diseases in patients with overt proteinuria who manifest the C allele, especially the homozygous CC phenotype.
Notes
Conflict of interest
All authors have no conflicts of interest to declare.
Funding
This work was supported by the Grant of HRH Princess Maha Chakri Sirindhorn Medical Center (contact number 435/2562).
Authors’ contributions
Conceptualization, Funding acquisition: CK
Data curation, Project administration: PP
Formal analysis: CK, SP, PP
Investigation: CK, PP, RY
Methodology: CK, PP
Visualization: CK, PP, TD
Writing–original draft: CK, PP, TP, KW, TD
Writing–review & editing: CK, TP, KW, TD
All authors read and approved the final manuscript.
Acknowledgements
The authors gratefully acknowledge use of the services and facilities of HRH Princess Maha Chakri Sirindhorn Medical Center (MSMC). We are deeply grateful for the grant supported by HRH Princess MSMC, Srinakharinwirot University, Thailand. We also would like to express our cordial thanks to Miss Punchita Rujirachaivej and her colleague in laboratory of molecular biology of MSMC for technical assistance, as well as Dr. Aaron Chow (Michigan State University) for critical reading of the manuscript. In addition, we thank Dr. Wittaya Jomoui (Department of Pathology, Srinakharinwirot University, Thailand) for the excellent information about molecular basis of diabetes and Assistant Professor Dr. Suchin Worawichawong (Department of Pathology, Mahidol University, Thailand) for the kidney pathology evaluation.